MRI recon server
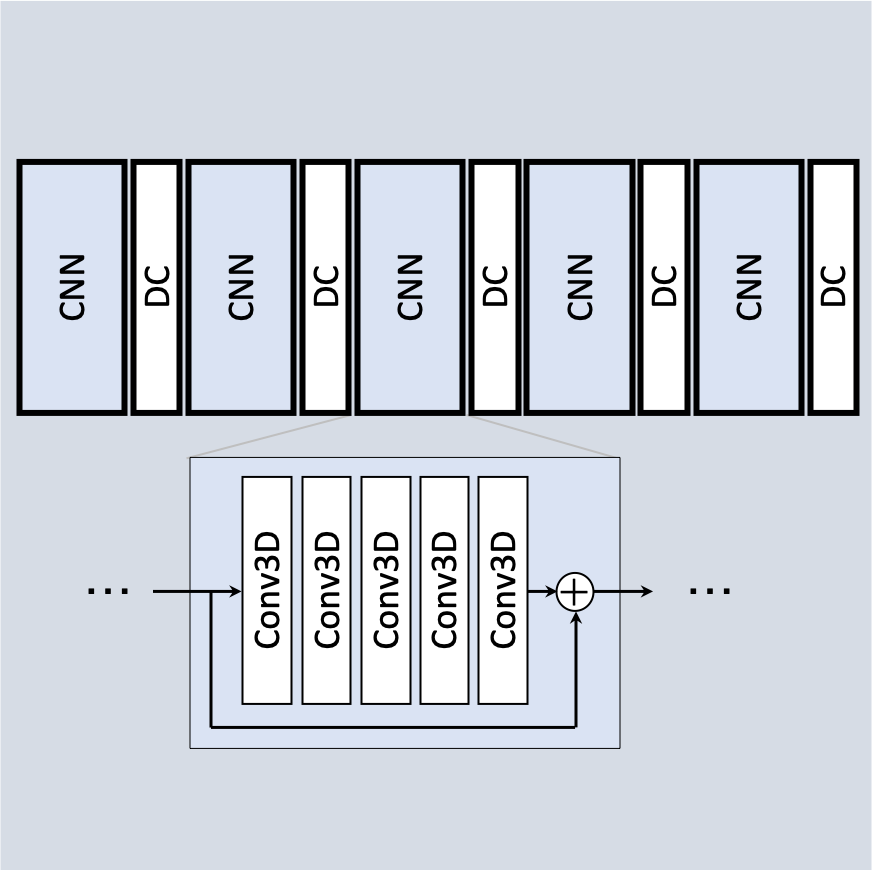
Integration and utilization of research MRI reconstruction methods can be hindered by computation lag and ease of use. An MRI reconstruction server was implemented using the Yarra framework, utilizing RIT‑HPC resources for computationally-intensive processes.
- rapid prototyping of new methods
- scalable resource allocation
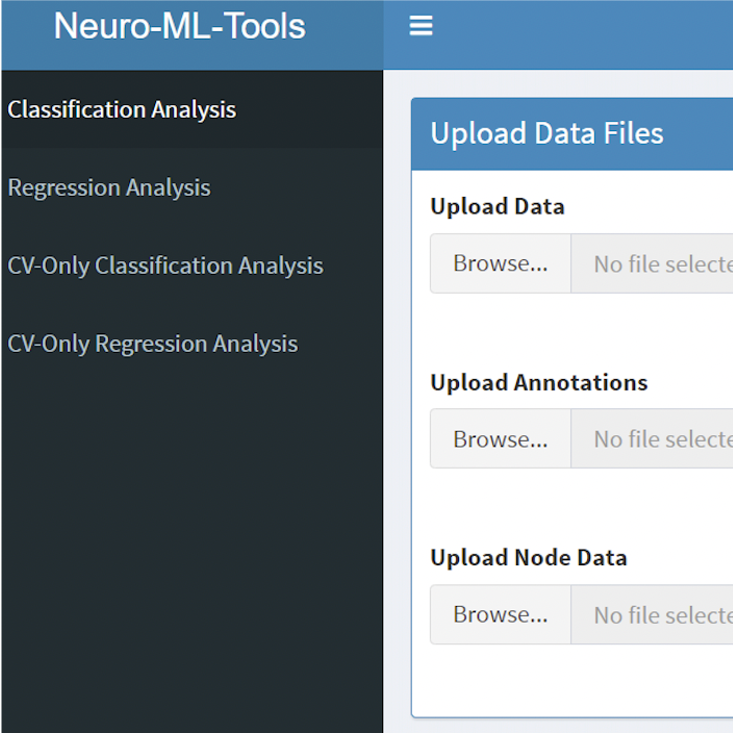
- ease of use for clinically-focused researchers
- automated end-to-end data transfer
- systematic archiving of research data and output
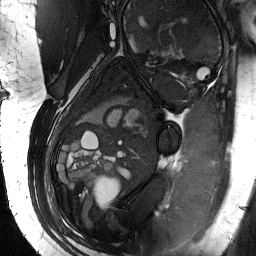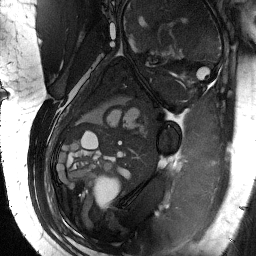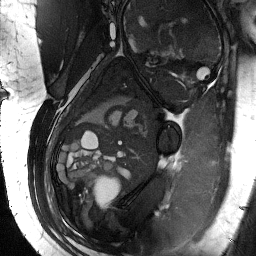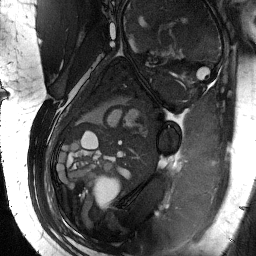
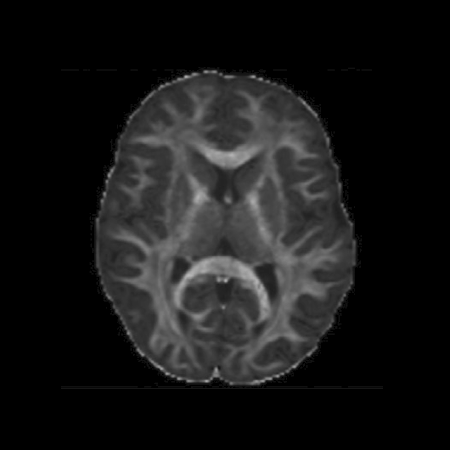
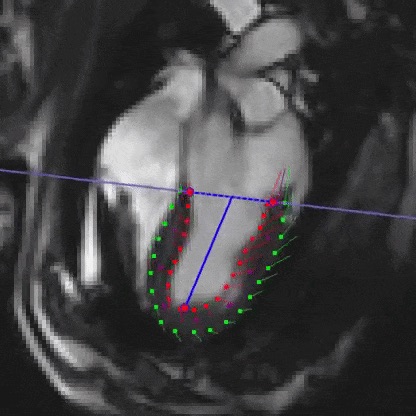
- enables computation of motion-corrected fetal cardiac MRI